Customizing and exporting graphical models and CPTs as image (pdf, png)
In [1]:
from pylab import *
import matplotlib.pyplot as plt
In [2]:
import pyagrum as gum
import pyagrum.lib.notebook as gnb
In [3]:
bn = gum.fastBN("a->b->c->d;b->e->d->f;g->c")
gnb.flow.row(bn, gnb.getInference(bn))
customizing colours and width for model and inference
In [4]:
def nodevalue(n):
return 0.5 if n in "aeiou" else 0.7
def arcvalue(a):
return (10 - a[0]) * a[1]
def arcvalue2(a):
return (a[0] + a[1] + 5) / 22
gnb.showBN(
bn,
nodeColor={n: nodevalue(n) for n in bn.names()},
arcWidth={a: arcvalue(a) for a in bn.arcs()},
arcLabel={a: f"v={arcvalue(a):02d}" for a in bn.arcs()},
arcColor={a: arcvalue2(a) for a in bn.arcs()},
)
In [5]:
gnb.showInference(
bn,
targets={"a", "g", "f", "b"},
evs={"e": 0},
nodeColor={n: nodevalue(n) for n in bn.names()},
arcWidth={a: arcvalue(a) for a in bn.arcs()},
)
In [6]:
gnb.flow.row(
gnb.getBN(bn, nodeColor={n: nodevalue(n) for n in bn.names()}, arcWidth={a: arcvalue(a) for a in bn.arcs()}),
gnb.getInference(bn, nodeColor={n: nodevalue(n) for n in bn.names()}, arcWidth={a: arcvalue(a) for a in bn.arcs()}),
)
In [7]:
mycmap = plt.get_cmap("Reds")
formyarcs = plt.get_cmap("winter")
gnb.flow.row(
gnb.getBN(
bn,
nodeColor={n: nodevalue(n) for n in bn.names()},
arcColor={a: arcvalue2(a) for a in bn.arcs()},
cmapNode=mycmap,
cmapArc=formyarcs,
),
gnb.getInference(
bn,
nodeColor={n: nodevalue(n) for n in bn.names()},
arcColor={a: arcvalue2(a) for a in bn.arcs()},
arcWidth={a: arcvalue(a) for a in bn.arcs()},
cmapNode=mycmap,
cmapArc=formyarcs,
),
)
Modifying graph’s layout
Every graph or graphical models can be translated into a pyDot’s representaton (a pydot.Dot object). In this graphical representation, it is possible to manipulate the positions of the node. pyAgrum proposes two functions gum.utils.dot_layout to help modifying this layout.
Layout for Bayesian network
In [8]:
import pyagrum as gum
import pyagrum.lib.notebook as gnb
import pyagrum.lib.utils as gutils
import pyagrum.lib.bn2graph as gumb2g
bn = gum.fastBN("A->B<-C<-D")
bn2 = gum.fastBN("A->B->C<-D")
graph = gumb2g.BN2dot(bn)
graph2 = gumb2g.BN2dot(bn2)
l = gutils.dot_layout(graph)
print(f"Layout proposed by dot for BN :{l}")
gutils.apply_dot_layout(graph2, l)
graph3 = gumb2g.BN2dot(bn2)
# l["C"],l["A"]=l["A"],l["C"]
l["D"], l["C"], l["B"], l["A"] = (
gutils.DotPoint(0, 0),
gutils.DotPoint(1, 1),
gutils.DotPoint(2, 2),
gutils.DotPoint(3, 3),
)
gutils.apply_dot_layout(graph3, l)
gnb.flow.row(bn, bn2, graph2, graph3, captions=["BN", "BN2", "BN2 with the same layoutas BN", "Layout changed by hand"])
Layout proposed by dot for BN :{'C': DotPoint(x=1.375, y=1.25), 'B': DotPoint(x=0.875, y=0.25), 'D': DotPoint(x=1.375, y=2.25), 'A': DotPoint(x=0.375, y=1.25)}
Layout for other graphical models and for inference
In [9]:
import pyagrum as gum
import pyagrum.lib.notebook as gnb
import pyagrum.lib.utils as gutils
import pyagrum.lib.id2graph as gum2gr
model = gum.fastID("*D->$L<-E<-H->L;E->D")
gnb.flow.add(model)
gum.config.push()
gum.config["influenceDiagram", "utility_shape"] = "diamond"
figure = gum2gr.ID2dot(model)
l = gutils.dot_layout(figure)
# changing LAYOUT
# making some horizontal space
for i, p in l.items():
l[i] = gutils.DotPoint(1.5 * p.x, p.y)
# E at the vertical of L, at the horizontal of D
l["E"] = gutils.DotPoint(l["L"].x, l["D"].y)
# H symetric of D w.r.t (EL)
l["H"] = gutils.DotPoint(2 * l["E"].x - l["D"].x, l["D"].y)
gutils.apply_dot_layout(figure, l)
gnb.flow.add(figure)
gnb.flow.display()
gum.config.pop()
In [10]:
import pyagrum as gum
import pyagrum.lib.notebook as gnb
import pyagrum.lib.utils as gutils
import pyagrum.lib.id2graph as gum2gr
model = gum.fastID("*D->$L<-E<-H->L;E->D")
gnb.flow.add(gnb.getInference(model))
figure = gum2gr.LIMIDinference2dot(model, evs={}, targets={}, size=None, engine=None)
l = gutils.dot_layout(figure)
# changing LAYOUT
# making some horizontal space
for i, p in l.items():
l[i] = gutils.DotPoint(3 * p.x, p.y)
# E at the vertical of L, at the horizontal of D
l["E"] = gutils.DotPoint(l["L"].x, l["D"].y)
l["D"] = gutils.DotPoint(l["D"].x / 2, l["D"].y)
l["H"] = gutils.DotPoint(l["L"].x * 3 / 2, l["D"].y)
gutils.apply_dot_layout(figure, l)
gnb.flow.add(figure)
gnb.flow.display()
Exporting model and inference as image
Exporting as image (pdf, png, etc.) has been gathered in 2 functions : pyagrum.lib.image.export() and pyagrum.lib.image.exportInference(). The argument are the same as for pyagrum.notebook.show{Model} and pyagrum.notebook.show{Inference}.
In [11]:
import pyagrum.lib.image as gumimage
from IPython.display import Image # to display the exported images
In [12]:
gumimage.export(bn, "out/test_export.png")
Image(filename="out/test_export.png")
Out[12]:

In [13]:
bn = gum.fastBN("a->b->d;a->c->d[3]->e;f->b")
gumimage.export(
bn,
"out/test_export.png",
nodeColor={"a": 1, "b": 0.3, "c": 0.4, "d": 0.1, "e": 0.2, "f": 0.5},
arcColor={(0, 1): 0.2, (1, 2): 0.5},
arcWidth={(0, 3): 0.4, (3, 2): 0.5, (2, 4): 0.6},
)
Image(filename="out/test_export.png")
Out[13]:

In [14]:
gumimage.exportInference(bn, "out/test_export.png")
Image(filename="out/test_export.png")
Out[14]:

In [15]:
gumimage.export(bn, "out/test_export.pdf")
Link to out/test_export.pdf
exporting inference with evidence
In [16]:
bn = gum.loadBN("res/alarm.dsl")
gumimage.exportInference(
bn,
"out/test_export.pdf",
evs={"CO": 1, "VENTLUNG": 1},
targets={
"VENTALV",
"CATECHOL",
"HR",
"MINVOLSET",
"ANAPHYLAXIS",
"STROKEVOLUME",
"ERRLOWOUTPUT",
"HBR",
"PULMEMBOLUS",
"HISTORY",
"BP",
"PRESS",
"CO",
},
size="15!",
)
Link to out/test_export.pdf
Other models
Other models can also use these functions.
In [17]:
infdiag = gum.loadID("res/OilWildcatter.bifxml")
gumimage.export(infdiag, "out/test_export.pdf")
Link to out/test_export.pdf
In [18]:
gumimage.exportInference(infdiag, "out/test_export.pdf")
Link to out/test_export.pdf
Exporting any object with toDot() method
In [19]:
import pyagrum.causal as csl
obs1 = gum.fastBN("Smoking->Cancer")
modele3 = csl.CausalModel(obs1, [("Genotype", ["Smoking", "Cancer"])], True)
gumimage.export(modele3, "out/test_export.png") # a causal model has a toDot method.
Image(filename="out/test_export.png")
Out[19]:

In [20]:
bn = gum.fastBN("a->b->c->d;b->e->d->f;g->c")
ie = gum.LazyPropagation(bn)
jt = ie.junctionTree()
gumimage.export(jt, "out/test_export.png") # a JunctionTree has a method jt.toDot()
Image(filename="out/test_export.png")
Out[20]:
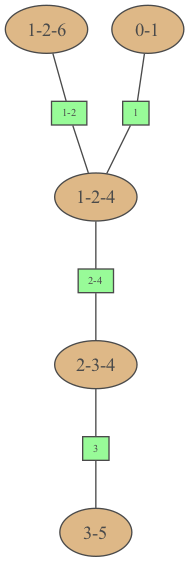
… or even a string in dot syntax
In [21]:
gumimage.export(
jt.toDotWithNames(bn), "out/test_export.png"
) # jt.toDotWithNames(bn) creates a dot-string for a junction tree with names of variables
Image(filename="out/test_export.png")
Out[21]:

Exporting to pyplot
In [22]:
import matplotlib.pyplot as plt
bn = gum.fastBN("A->B->C<-D")
plt.imshow(gumimage.export(bn))
plt.show()
plt.imshow(gumimage.exportInference(bn, size="15!"))
plt.show()
plt.figure(figsize=(10, 10))
plt.imshow(gumimage.exportInference(bn, size="15!"))
plt.show()
Exporting CPTs, sideBySide, explain.Information (and other html strings)
pyAgrum uses the package playwright in order to export html string. It proposes a function pyagrum.utils.async_html2image. In pyagrum.notebook, the functions get... return HTML objects. The functions show... display the result.
As a result, every pyagrum.lib.notebook.get... can be exported as pdf, png … using pyagrum.utils.async_html2image.
In [23]:
bn = gum.fastBN("A->B<-C", 3)
await gutils.async_html2image(gnb.getTensor(bn.cpt("B")), "out/cpt_B_.pdf")
In [24]:
await gutils.async_html2image(gnb.getSideBySide(bn, bn.cpt("A"), bn.cpt("B"), bn.cpt("C")), "out/sideBySide.pdf")
In [25]:
import pyagrum.explain as gexplain
await gutils.async_html2image(gexplain.getInformation(bn), "out/informationBN.pdf")
In [ ]:

