pyagrum.lib.explain
The purpose of pyagrum.lib.explain is to give tools to explain and interpret the structure and parameters of a Bayesian network.
Dealing with independence
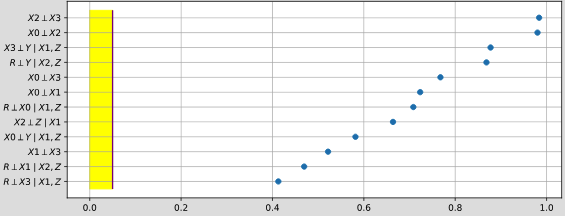
- pyagrum.lib.explain.independenceListForPairs(bn, filename, target=None, plot=True, alphabetic=False)
get the p-values of the chi2 test of a (as simple as possible) independence proposition for every non arc.
- Parameters:
bn (gum.BayesNet) – the Bayesian network
filename (str) – the name of the csv database
alphabetic (bool) – if True, the list is alphabetically sorted else it is sorted by the p-value
target ((optional) str or int) – the name or id of the target variable
plot (bool) – if True, plot the result
- Returns:
the list
Dealing with mutual information and entropy

- pyagrum.lib.explain.getInformation(bn, evs=None, size=None, cmap=<matplotlib.colors.LinearSegmentedColormap object>)
get a HTML string for a bn annotated with results from inference : entropy and mutual information
- Parameters:
bn (pyagrum.BayesNet) – the model
evs (Dict[str|int,str|int|List[float]]) – the observations
size (int|str) – size of the rendered graph
cmap (matplotlib.colours.Colormap) – the cmap
- Returns:
return the HTML string
- Return type:
str
- pyagrum.lib.explain.showInformation(bn, evs=None, size=None, cmap=<matplotlib.colors.LinearSegmentedColormap object>)
diplay a bn annotated with results from inference : entropy and mutual information
- Parameters:
bn (pyagrum.BayesNet) – the model
evs (Dict[str|int,str|int|List[float]]) – the observations
size (int|str) – size of the rendered graph
cmap (matplotlib.colours.Colormap) – the cmap
Dealing with ShapValues
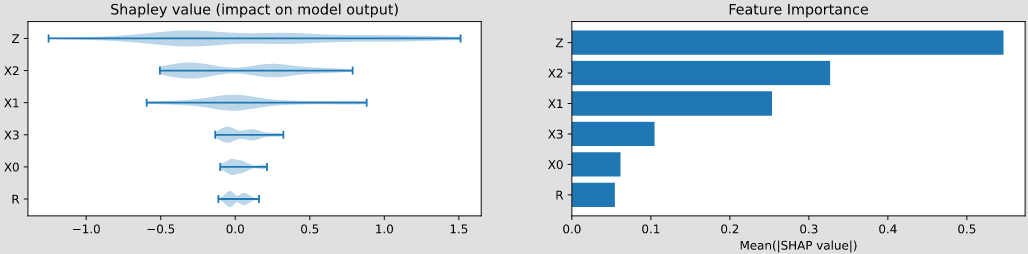
- class pyagrum.lib.explain.ShapValues(bn, target, logit=True)
Bases:
objectClass to compute Shapley values for a target variable in a Bayesian network.
- causal(df, y=1, sample_size=200, plot=False, plot_importance=False, percentage=False, filename=None)
Computes the causal Shapley values for each variable.
Parameters:
- dfpandas DataFrame
The input data for which to compute the Shapley values.
- yint, optional
The target class for which to compute the Shapley values (default is 1).
- sample_sizeint, optional
The number of samples to use for the background data (default is 200).
- plotbool, optional
If True, plots the waterfall or beeswarm plot depending on the number of rows in df (default is False).
- plot_importancebool, optional
If True, plots the bar chart of feature importance (default is False).
- percentage: bool
if True, the importance plot is shown in percent.
- filenamestr, optional
If provided, saves the plots to the specified filename instead of displaying them.
Returns:
: Dict[str, float]
A dictionary containing the importances of each variable in the input data.
- type percentage:
bool- param percentage:
- type filename:
str- param filename:
- conditional(df, y=1, plot=False, plot_importance=False, percentage=False, filename=None)
Computes the conditional Shapley values for each variable.
Parameters:
- dfpandas DataFrame
The input data for which to compute the Shapley values.
- yint, optional
The target class for which to compute the Shapley values (default is 1).
- plotbool, optional
If True, plots the waterfall or beeswarm plot depending on the number of rows in df (default is False).
- plot_importancebool, optional
If True, plots the bar chart of feature importance (default is False).
- percentage: bool
if True, the importance plot is shown in percent.
- filenamestr, optional
If provided, saves the plots to the specified filename instead of displaying them.
Returns:
: Dict[str, float]
A dictionary containing the importances of each variable in the input data.
- type y:
int- param y:
- type plot:
bool- param plot:
- type plot_importance:
bool- param plot_importance:
- type percentage:
bool- param percentage:
- type filename:
str- param filename:
- marginal(df, y=1, sample_size=200, plot=False, plot_importance=False, percentage=False, filename=None)
Computes the marginal Shapley values for each variable.
Parameters:
- dfpandas DataFrame
The input data for which to compute the Shapley values.
- yint, optional
The target class for which to compute the Shapley values (default is 1).
- sample_sizeint, optional
The number of samples to use for the background data (default is 200).
- plotbool, optional
If True, plots the waterfall or beeswarm plot depending on the number of rows in df (default is False).
- plot_importancebool, optional
If True, plots the bar chart of feature importance (default is False).
- percentage: bool
if True, the importance plot is shown in percent.
Returns:
: Dict[str, float]
A dictionary containing the importances of each variable in the input data.
- type percentage:
bool- param percentage:
- type filename:
str- param filename:
Dealing with generalized Markov Blankets
A structural property of Bayesian networks is the Markov boundary of a node. A Markov blanket of a node is a set of nodes that renders the node independent of all other nodes in the network. The Markov boundary is the closest Markov blanket. A Markov boundary of a node is composed of its parents, its children, and the parents of its children. More generally, one can define the generalized \(k\)-Markov blanket of a node as the union of the markov blanket of the nodes of its \((k-1)\)-Markov blanket. So, if a node belongs to the \(k\)-Markov blanket of the node \(X\), \(k\) is a kind of measure of its proximity to \(X\).
